From the Field
Genevieve Housman was awarded a Leakey Foundation Research Grant during our fall 2015 cycle for her project entitled “Assessment of DNA methylation patterns in primate skeletal tissues.” In March we featured her work on our blog, which you can read by clicking here. Below she updates us on her progress.

As I wrap up a significant portion of my data collection, I feel it’s time to send out some news from the field.
Being a molecular anthropologist, I don’t exactly have a “field site.” Rather, my work takes place in the clean, safe, and temperature controlled environments of laboratories. In particular, I have collected and processed data at both the Molecular Anthropology Laboratory at Arizona State University and the Genetics Department at the Texas Biomedical Research Institute (TxBiomed).
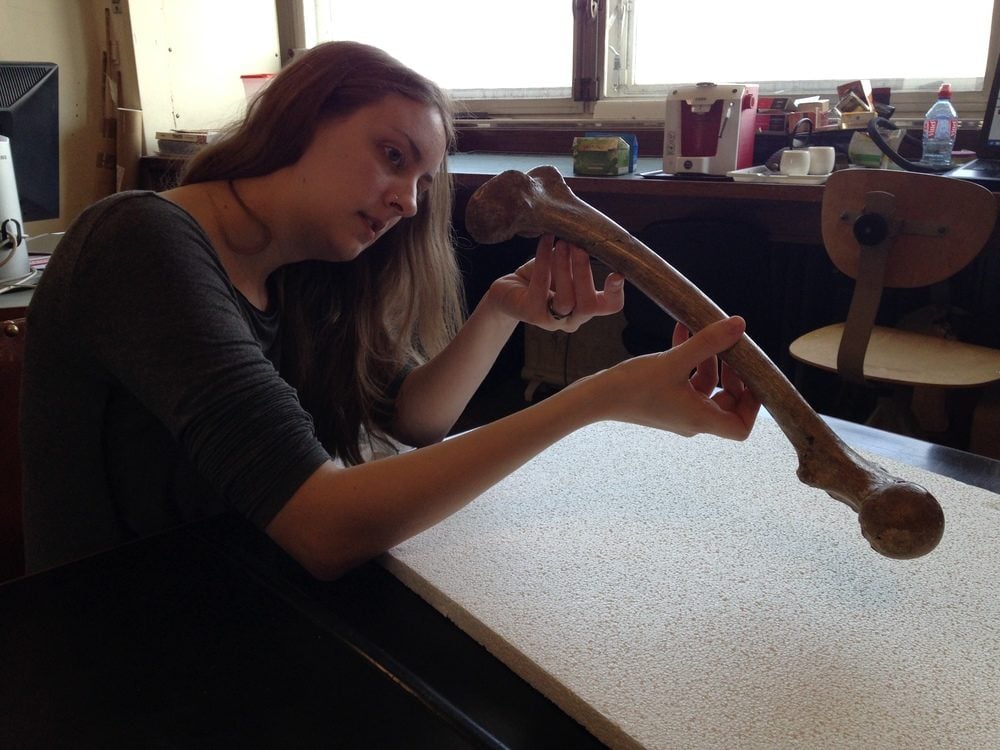
My research centers on understanding how epigenetic changes contribute to complex skeletal phenotypes in non-human primates. Specifically, I study how variations in DNA methylation patterns relate to variations in femur bone morphologies and the development of skeletal disorders such as osteoarthritis.
To do this, I collected morphological measurements and tissue samples from the femora of over 300 non-human primate specimens. These included mostly baboons, as well as chimpanzees, macaques, marmosets, and vervets. I did this initial collection at TxBiomed one year ago. It involved dissecting out previously necropsied and frozen limbs, collecting 29 linear measurements and 55 digital landmarks from each specimen, and sampling cartilage scrapings and drilled bone cores from each specimen. All together, these efforts produced over 25,000 morphological data points and over 500 tissue specimens.
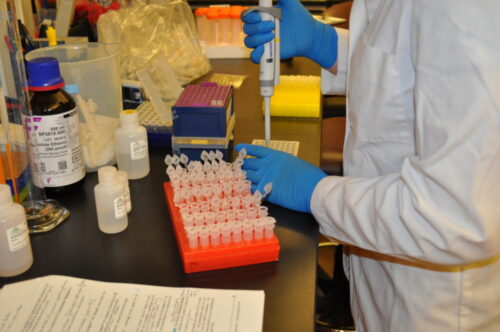
My focus then shifted towards processing and extracting DNA from my tissue specimens so that they could be used in downstream efforts to collect DNA methylation data at a genome-wide level using an advanced probe-based array technology and at a gene-specific level using bisulfite sequencing methods.
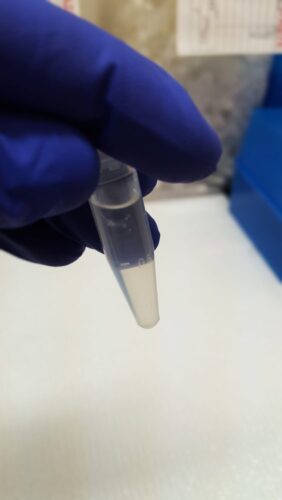
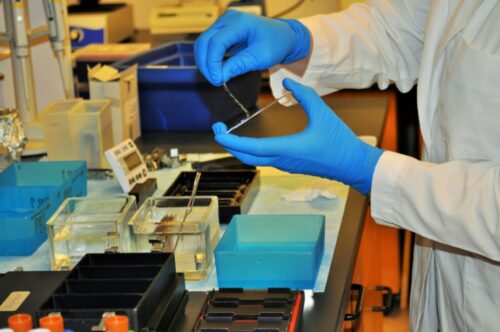
First, I pulverized my bone and cartilage samples so that DNA could be more readily extracted from them. To avoid contamination, this process required a series of equipment cleaning steps in between each sample. Then, each sample was extracted using a phenol-chloroform protocol, and DNA concentrations were quantified.
With these steps completed, I just recently began my downstream genome-wide DNA methylation work. This summer I again travelled to TxBiomed, where I assembled and ran 160 of my DNA extractions on probe-based arrays. These arrays examine the DNA methylation levels of over 850,000 loci throughout the genome, and the data look good so far!
I am currently working on statistically comparing these DNA methylation patterns across my different species and with respect to my skeletal phenoypes, and I will begin my gene-specific data collection soon. Hopefully, I’ll have exciting results to share with everyone at the upcoming meetings this year!